Sequana documentation#
0.22
SEQUANA#
)




- How to cite:
Citations are important for us to carry on developments. For Sequana library (including the pipelines), please use
Cokelaer et al, (2017), 'Sequana': a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
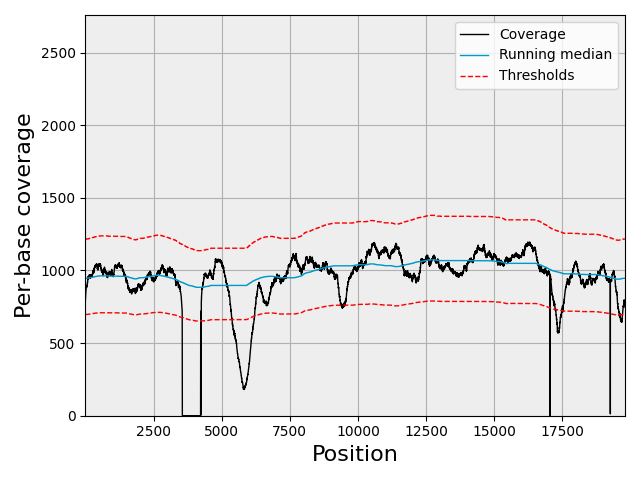
For the genome coverage tool (sequana_coverage): Desvillechabrol et al, 2018: detection and characterization of genomic variations using running median and mixture models. GigaScience, 7(12), 2018. https://doi.org/10.1093/gigascience/giy110
For Sequanix: Desvillechabrol et al. Sequanix: A Dynamic Graphical Interface for Snakemake Workflows Bioinformatics, bty034, https://doi.org/10.1093/bioinformatics/bty034 Also available on bioRxiv (DOI: https://doi.org/10.1101/162701)
🔧 Overview and Installation#
Sequana is a Python library dedicated to bioinformatics. It is also a project that includes a set of pipelines related to NGS (new generation sequencing) including quality control, variant calling, coverage, taxonomy, transcriptomics. We also ship Sequanix, a graphical user interface for Snakemake pipelines.
Contributors#
Maintaining Sequana would not have been possible without users and contributors. Each contribution has been an encouragement to pursue this project. Thanks to all:
Changelog :memo:#
Version |
Description |
|---|---|
0.22.0 |
|
0.21.2 |
|
0.21.1 |
|
0.21.0 |
|
0.20.0 |
|
0.19.6 |
|
0.19.5 |
|
0.19.4 |
|
0.19.3 |
|
0.19.2 |
|
0.19.1 |
|
0.19.0 |
|
0.18.0 |
|
0.17.3 |
|
0.17.2 |
|
0.17.1 |
|
0.17.0 |
|
0.16.9 |
|
0.16.8 |
|
0.16.7 |
|
0.16.6 |
|
0.16.5 |
|
0.16.4 |
|
0.16.3 |
|
0.16.2 |
|
0.16.1 |
|
0.16.0 |
|
0.15.4 |
|
0.15.3 |
|
0.15.2 |
|
0.15.1 |
|
0.15.0 |
|
0.14.6 |
|
0.14.5 |
|
0.14.4 |
|
0.14.3 |
|
0.14.2 |
|
0.14.1 |
|
0.14.0 |
|
0.13.X |
|
0.12.X |
|
Any :question: Feel free to [open an issue](sequana/sequana#issues)
What is Sequana?#
Sequana is a versatile bioinformatics tool that provides:
A Python library dedicated to NGS analysis (file-format wrappers, plots, coverage, taxonomy, enrichment, RNA-seq, …).
A family of pipelines based on Snakemake, each living in its own GitHub repository.
- A set of standalone applications:
sequana_coverage — genome coverage analysis with confidence intervals.
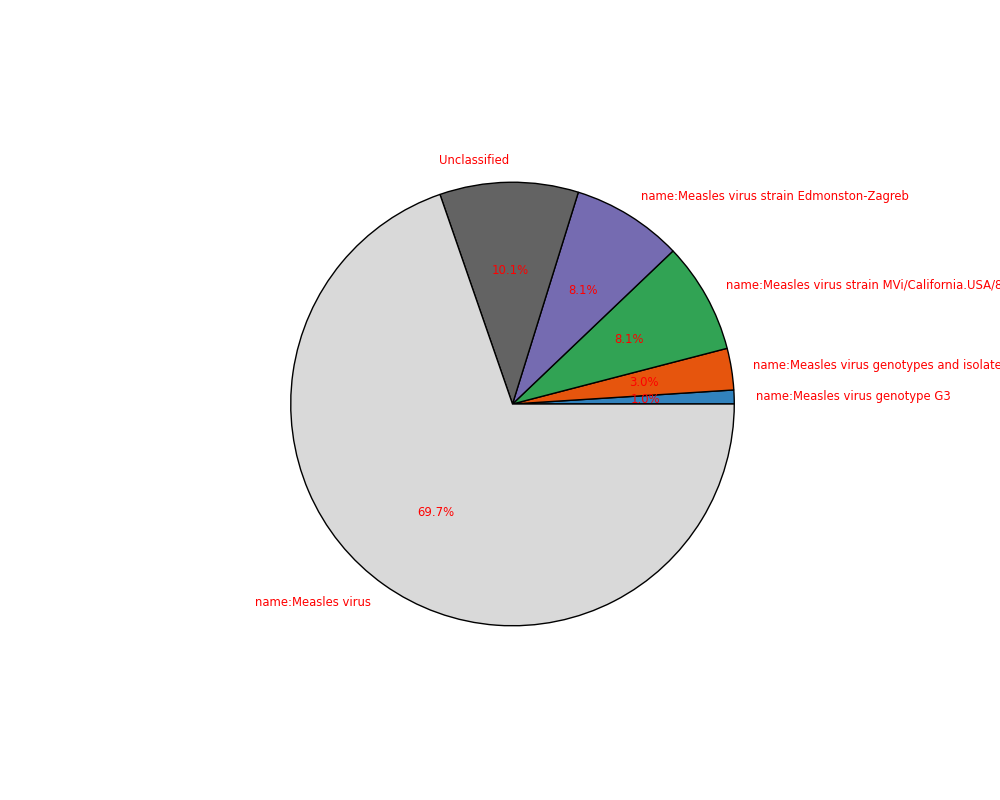
sequana_taxonomy — quick taxonomy on a FastQ file (Kraken + Krona).
~30 other utility sub-commands grouped under the top-level
sequanaCLI (see Applications (standalone)).
Pipelines cover NGS quality control, variant calling, coverage analysis, taxonomy, de-novo assembly, RNA-seq, variant calling and more — see the Pipelines catalogue.
Sequana can be used either as a Python library (developers, library users) or via its pipelines and standalones (end users). To join the project, please let us know on github.
Installation
pip install sequana
Examples
Visit our example gallery to use the Python library
NGS pipelines
Browse the Snakemake pipelines
Standalone applications
The sequana CLI and its sub-commands
Pipeline users
Library users
Reference & developers
- Developer guide
- Applications (standalone)
- Sequanix Tutorial
- Wrappers
- bowtie2/align
- bowtie2/build
- add_read_group
- bam_coverage
- bcl2fastq
- blast
- busco
- bz2_to_gz
- canu
- consensus
- digital_normalisation
- dsrc_to_gz
- falco
- fastp
- fastq_stats
- fastqc
- feature_counts
- freebayes
- freebayes_vcf_filter
- gz_to_bz2
- hmmbuild
- hmmscan
- index
- longorfs
- macs3
- makeblastdb
- mark_duplicates
- minimap2
- multiqc
- polypolish
- predict
- prokka
- quast
- rulegraph
- sambamba_filter
- sambamba_markdup
- samtools_depth
- sequana_coverage
- sequana_taxonomy
- snpeff
- snpeff_add_locus_in_fasta
- sort
- spades
- trinity
- trinity_quantify
- unicycler
- API reference
- Glossary